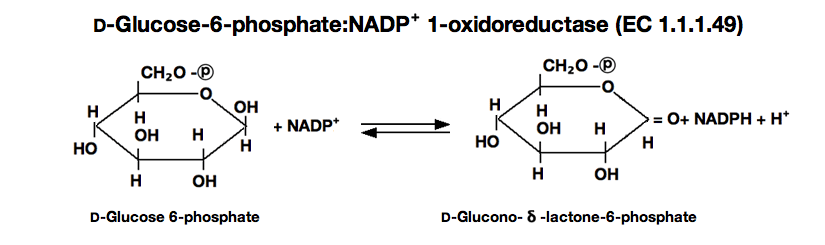
GLUCOSE-6-PHOSPHATE DEHYDROGENASE from Microorganism
G6D-321
| Appearance: | White amorphous powder, lyophilized | ||
|---|---|---|---|
| Activity: | GradeⅢ 200U/mg-solid or more |
||
| Contaminants: | Creatine phosphokinase ≤1×10⁻³% Phosphoglucomutase ≤1×10⁻³% 6-Phosphogluconate dehydrogenase ≤5×10⁻³% Phosphoglucose isomerase ≤1×10⁻²% Glutathione reductase ≤1×10⁻³% Hexokinase ≤1×10⁻²% Myokinase ≤1×10⁻²% NADH oxidase ≤1×10⁻²% NADPH oxidase ≤1×10⁻²% |
||
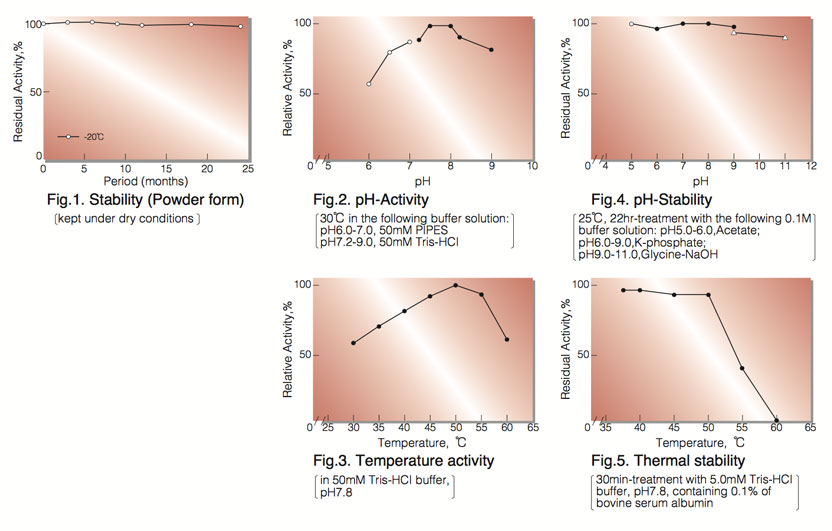
| Stability: | Stable at -20°C for at least One year (Fig.1) |
|---|---|
| Molecular weight: | approx. 140,000 (by gel filtration) |
| Michaelis constants: | NAD⁺ linked 2.4×10⁻⁴M (NAD⁺), 4.7×10⁻⁴M (G-6-P) NADP⁺ linked 7.4×10⁻⁶M (NADP⁺), 3.2×10⁻⁴M (G-6-P) |
| Inhibitors: | Metal ions, iodoacetamimide, SDS etc. |
| Optimum pH : | 7.8(Fig.2) |
| Optimum temperature: | 50°C-55°C(Fig.3) |
| pH Stability: | pH 5.0-11.0 (25°C, 22hr)(Fig.4) |
| Thermal stability: | below 50°C (pH 7.8, 30min)(Fig.5) |
| Substrate specificty: | (Table 1) |
| Effect of various chemicals: | (Table 2) |
APPLICATIONS
The enzyme is useful for enzymatic determination of NAD⁺(NADP⁺) and G-6-P, and activities of phosphoglucose isomerase, phosphoglucomutase and hexokinase. The enzyme is also used for enzymatic determination of glucose and creatine phosphokinase activity when coupled with hexokinase (HXK-311).
ASSAY
Principle:
glucose-6-phophate dehydrogenase(G-6-PDH)
D-Glucose-6-phosphate(G-6-P)+NAD⁺ ►
D-Glucono-δ-lactone-6-phosphate+NADH+H⁺
The appearance of NADH is measured at 340nm by spectrophotometry.
Unit definition:
One unit causes the formation of one micromole of NADH per minute under the conditions described below.
Method:
| A. Tris-HCl buffer, pH 7.8: | 55mM (contaning 3.3mM magnesium chloride) |
|---|---|
| B. NAD⁺ solution: | 60mM (Should be prepared fresh) |
| C. G-6-P solution: | 0.1M D-glucose-6-phosphate (Should be prepared fresh) |
| D. Enzyme diluent: | 5mM Tris-HCl buffer, pH 7.5, containing 0.1% of bovine serum albumin. |
Procedure
| Concentration in assay mixture | |
|---|---|
| Tris-HCl buffer | 50 mM |
| G-6-P | 3.3 mM |
| NAD⁺ | 2.0 mM |
| MgCl₂ |
3.0 mM |
| BSA | 33µg/ml |
1. Prepare the following reaction mixture in a cuvette (d=1.0cm) and equilibrate at 30°C for about 5 minutes.
2.7 ml Tris-HCl buffer, pH 7.8 (A)
0.1 ml NAD⁺ solution (B)
0.1 ml G-6-P solution, pH 7.8 (C)
2. Add 0.1ml of the enzyme solution *and mix by gentle inversion.
3. Record the increase in optical density at 340nm against water for 4 to 5 minutes in a spectrophotometer
thermostated at 30°C, and calculate the ΔOD per minute from the initial linear portion of the curve (ΔOD test).
At the same time, measure the blank rate (ΔOD blank) by using the same method as the test except that the enzyme diluent(D) is added instead of the enzyme solution.
* Dissolve the enzyme preparation in ice-cold enzyme diluent (D) and dilute to 0.05-0.20U/ml with the same buffer, immediately before the assay.
Calculation
Activity can be calculated by using the following formula :

ΔOD/min (OD test−OD blank ) ×Vt × df
Volume activity (U/ml) = =ΔOD/min×4.82×df
6.22×1.0×Vs
Weight activity (U/mg)=(U/ml)×1/C
- Vt
- : Total volume (3.0ml)
- Vs
- : Sample volume (0.1ml)
- 6.22
- : Millimolar extinction coefficient of NADH under the assay condition (㎠/micromole)
- 1.0
- : Light path length (cm)
- df
- : Dilution factor
- C
- : Enzyme concentration in dissolution
Table 1. Substrate Specificity of Glucose-6-phosphate dehydrogenase
[3.3mM of substrate, 50mM Tris-HCl buffer, pH 7.8]
| Substrate | Relative activity(%) |
|---|---|
| Glucose-6-phosphate | 100 |
| Fructose-6-phosphate | 0 |
| Glucose-1-phosphate | 0 |
| Gluconate-6-phosphate | 0 |
| Chemical | Concn.(mM) | Residual activity(%) |
Chemical | Concn.(mM) | Residual activity(%) |
|---|---|---|---|---|---|
| None | − | 100 | NEM | 2.0 | 94 |
| Metal salt | 2.0 | PCMB | 2.0 | 103 | |
| AgNO₃ |
74 | MIA |
2.0 | 99 |
|
| Ba(OAc)₂ | 100 | Iodoacetamide | 2.0 | 0 | |
| CaCl₂ | 95 | EDTA | 5.0 | 102 | |
| Cd(OAc)₂ | 84 | (NH₄)₂SO₄ | 20.0 | 99 | |
| CoCl₂ | 94 | Borate | 20.0 | 98 | |
| CuSO₄ | 84 | o-Phenanthroline | 2.0 | 100 | |
| FeCl₃ | 0 |
α,α′-Dipyridy | 2.0 | 102 | |
| FeSO₄ | 0 | Urea | 2.0 | 99 | |
| HgCl₂ | 87 | Guanidine | 2.0 | 100 | |
| MgCl₂ | 100 | Hydroxylamine | 2.0 | 100 | |
| MnCl₂ | 97 | Na-cholate | 1.0% | 106 | |
| NiCl₂ | 94 | Triton X-100 | 1.0% | 104 | |
| Pb(OAc)₂ |
31 | Brij 35 | 1.0% | 12 | |
| Zn(OAc)₂ | 63 |
SDS | 0.1% | 1 | |
| ZnSO₄ | 76 | Tween 20 | 0.1% | 102 | |
| KF |
2.0 | 103 | Span 20 | 0.1% | 101 |
| NaF |
20.0 | 99 | DAC | 0.1% | 1 |
| NaN₃ | 20.0 | 102 |
Ac, CH₃CO; NEM, N-Ethylmaleimide; PCMB, p-Chloromercuribenzoate; MIA, Monoiodoacetate; EDTA, Ethylenediaminetetraacetate; SDS, Sodium dodecyl sulfate; DAC, Dimethylbenzylalkylammoniumchloride.