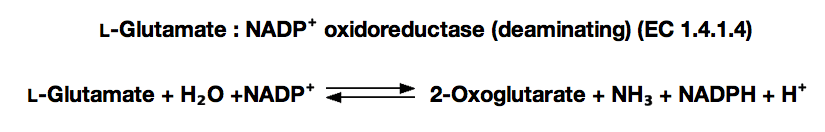
GLUTAMATE DEHYDROGENASE (NADP-dependent) from Proteus sp.
GTD-209, 309
| Appearance: | Solution with 50mM Tris-HCl buffer containing 0.05% NaN₃ and 5.0mM EDTA, pH 7.8 | ||
|---|---|---|---|
| Activity: | GradeⅡ∙Ⅲ 300U/mg-protein or more (9,000U/ml or more) |
||
| Contaminants: | NADPH oxidase ≤1.0×10⁻²% Glutathione reductase ≤1.0×10⁻²% (GradeⅡ-209) ≤1.0×10⁻¹% (GradeⅢ-309) |
||
| Stabilizer: | Ethylenediaminetetraacetic acid (EDTA) | ||
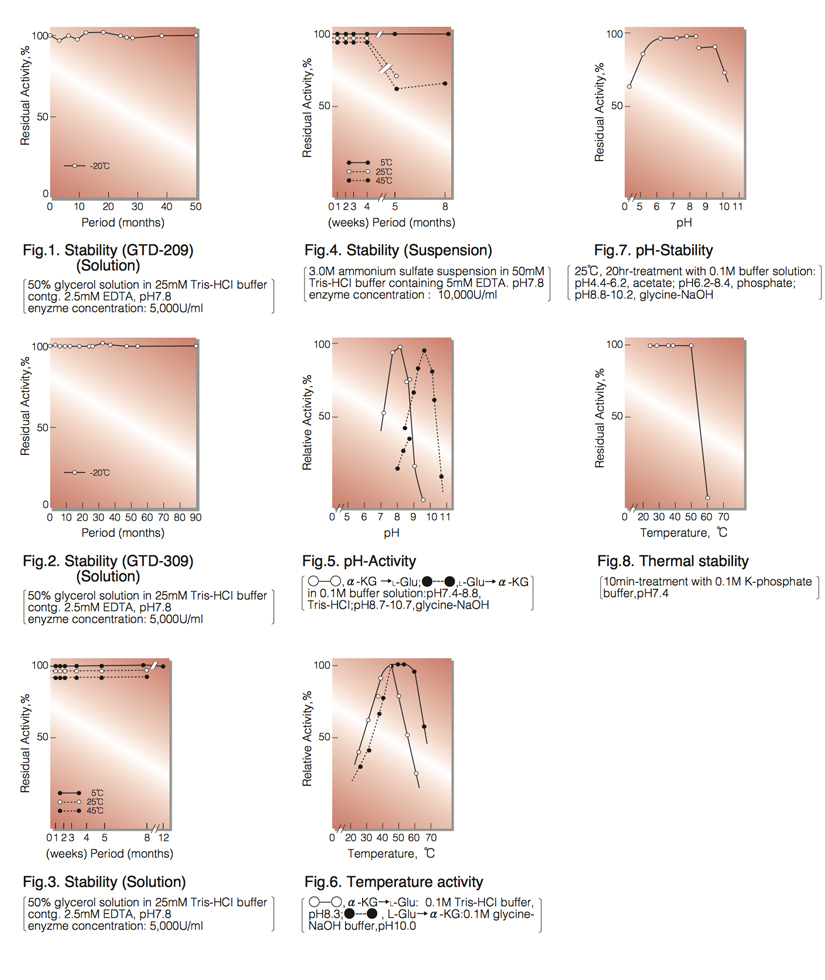
| Stability: | Stable at -20°C for at least One year (Fig.1,2) |
|---|---|
| Molecular weight: | approx. 300,000 |
| Isoelectric point : | 4.6 |
| Michaelis constants : | 1.1×10⁻³M (NH₃), 3.4×10⁻⁴M (α-Ketoglutarate) 1.2×10⁻³M (L-Glutamate), 1.4×10⁻⁵M (NADPH), 1.5×10⁻⁵M (NADP⁺) |
| Structure : | 6 subunits (M.W.50,000) per enzyme molecule |
| Inhibitors : | Hg⁺⁺, Cd⁺⁺, p-chloromercuribenzoate, pyridine, 4-4'-dithiopyridine, 2,2'-dithiopyridine |
| Optimum pH : | 8.5 (α-KG→L-Glu) 9.8 (L-Glu→α-KG)(Fig.5) |
| Optimum temperature: | 45°C (α-KG→L-Glu) 45-55°C (L-Glu→α-KG)(Fig.6) |
| pH Stability: | pH 6.0-8.5 (25°C, 20hr)(Fig.7) |
| Thermal stability: | below 50°C (pH 7.4, 10min)(Fig.8) |
| Substrate specificity: | (Table 1) |
APPLICATIONS
This enzyme is useful for enzymatic determination of NH₃, α-ketoglutaric acid and L-glutamic acid, and for assay of leucine aminopeptidase and urease. This enzyme is also used for enzymatic determination of urea when coupled with urease (URH-201) in clinical analysis.
ASSAY
Principle:
glutamate dehydrogenase
α-Ketoglutarate+NH₃+ NADPH + H⁺ ► L-Glutamate+NADP⁺+H₂O
The disappearance of NADPH is measured at 340nm by spectrophotometry.
Unit definition:
One unit causes the oxidation of one micromole of NADPH per minute under the conditions described below.
Method:
| A. Buffer solution: | 0.1M Tris-HCl buffer, pH 8.3 |
|---|---|
| B. NH₄Cl solution: | 3.3M |
| C. α-Ketoglutarate solution: | 0.225M (adjust the pH to 7.0-9.0 with NaOH)(Should be prepared fresh) |
| D. NADPH solution: | 7.5mM (Should be prepared fresh) |
| E. Enzyme diluent: | 50mM K-Phosphate buffer, pH 6.6 containing 0.2% BSA and 50mM EDTA |
Procedure
| Concentration in assay mixture | |
|---|---|
| Tris-HCl buffer | 85 mM |
| α-Ketoglutarate | 7.6 mM |
| NH₄Cl | 0.22 M |
| NADPH |
0.25mM |
| EDTA | 0.85mM |
1. Prepare the following reaction mixture in a cuvette (d=1.0cm) and equilibrate at 30°C for about 5 minutes.
2.5 ml Buffer solution (A)
0.2ml NH₄Cl solution (B)
0.1ml α-Ketoglutarate solution (C)
0.1ml NADPH solution (D)
2. Add 0.05ml of the enzyme solution* and mix by gentle inversion.
3. Record the decrease in optical density at 340nm against water for 2 to 3 minutes in a spectro-photometer
thermostated at 30°C and calculate the ΔOD per minute from the linear portion of the curve (ΔOD test).
At the same time, measure the blank rate (ΔOD blank) by using the same method as the test except that
the enzyme diluent (E) is added instead of the enzyme solution.
* Dilute the enzyme preparation to 0.4-0.9U/ml with ice-cold enzyme diluent (E), immediately before the assay.
Calculation
Activity can be calculated by using the following formula :

ΔOD/min (ΔOD test−ΔOD blank ) ×Vt × df
Volume activity (U/ml) = =ΔOD/min×9.486×df
6.22×1.0×Vs
Weight activity (U/mg)=(U/ml)×1/C
- Vt
- : Total volume (2.95ml)
- Vs
- : Sample volume (0.05ml)
- 6.22
- : Millimolar extinction coefficient of NADPH (㎠/micromole)
- 1.0
- : Light path length (cm)
- df
- : Dilution factor
REFERENCES
- H.Shimizu, T.Kuratsu and F.Hirata; J.Ferment.Technol., 57, 428 (1979).
| Substrate (50mM) | Relative activity(%) | Substrate (50mM) | Relative activity(%) |
|---|---|---|---|
| L-Glutamate | 100 | L-Glutamine | < 0.01 |
| L-Norvaline | 0.39 |
L-Aspartate | < 0.01 |
| L-α-Aminobutyrate | 0.19 | L-Asparagine | < 0.01 |
| L-Norleucine | 0.04 | L-Valine | < 0.01 |
| D,L-Homocysteine | 0.03 | L-Leucine | < 0.01 |
| L-Isoleucine | 0.02 | L-Alanine | < 0.01 |
| L-Methionine | < 0.01 |
Glutamate dehydrogenase:18U/ml of 0.1M glycine-NaOH buffer, pH 9.0 NADP⁺: 0.3mM